 |
|||||||||||||
|
 |
|||||||||||||
|
Project in Google Docs
Post traumatic stress disorder is an anxiety disorder that affects people of many different demographics. Post traumatic stress disorder has severe consequences in social functioning to those afflicted with the disorder and also often gives rise to comorbidity. Previous studies have shown that some people may be more susceptible to develop the disorder based on their genotype. The purpose of this project is to find a way to test for the presence of an allele which serves as a marker for predisposition to develop Post Traumatic Stress Disorder (PTSD) of which the entire etiology is not known. The presence of this allele represents only a small piece of information in a highly complex web of neurobiological function. This allele is present in many adults, some are homozygous for the allele and others are heterozygous L/L or S/S or S/L. They will be herein referred to as the Long or Short allele. The development of this test is crucial in preventing onset of the disease by reducing exposure to traumatic events like combat for example. Genetic testing is expensive and a more cost effective way to detect vulnerability is needed. A test that requires minimal training to perform and with materials that are cost-effective would be better than lengthy existing methods. Is there a way that we can quickly test a person's genotype without the cumbersome, expensive burden of polymerase chain reaction and gel electrophoresis? In this study, we aim to use some known techniques to streamline genetic testing making it more accessible. In order to accomplish this, we first used known methods of genotyping such as PCR and gel electrophoresis to establish a basis for genotype on n=3 samples (Mellman et al. 1996, Gerlerntner et al. 1993). One sample derived from the human eye cell line ARPE19 and 2 from volunteer blood donors. After successfully amplifying the region of interest from the sequence, we then sequenced those reactions to confirm our results from gel electrophoresis. We can use PCR and gel electrophoresis as a way to test our results we get with the new method. What is the most cost effective, simple way to test for allelic variation? In order to improve upon previous work, we plan to use a less invasive approach to extract and scan DNA samples: Urinalysis. This method of DNA extraction in combination with using molecular probes and paper chromatography, may provide a way to "quick test" for allelic variation.
RESULTS: NOT YET DETERMINED
CONCLUSION: NOT YET DETERMINED
Proposal for the rest of the project
(There is a kit from Qiagen which claims to extract an average 25ug of DNA from urine samples. We can first utilize this kit and later develop our own protocol to quickly extract DNA from urine samples.) Due to the 44bp deletion in the short version of the allele, we will use a generated probe that will hybridize to the region of interest located in the long version of the allele.
An additional probe that will hybridize to the sequences surrounding the deletion on the short version both will be different colors
The probe should be designed in a way that it can recognize the sequence, incubation/hybridization period is short and visualization of the results can be seen without the use of special equipment or machines.
In designing the probe, I would like to combine a variety of approaches to be able to detect hybridization of the probe and then visualize the results.
Paper chromatography, as used in the fields of chemistry as well as molecular biology is useful for quick visualization to detect the presence of a molecule/compound using reactive dyes that simply use nitrocellulose paper along with chemical tagging to produce a visual reaction(Zhang et al. 2000, Rule et al. 1996).
Using known genotypes positive for the long allele as a control, we should conduct testing simultaneously using the probe and paper chromatograph.
We can quantify our results by using a method similar to statistical analysis of Western Blotting. (statistical analysis)
SERT SEQUENCE

Heils et al. 1996
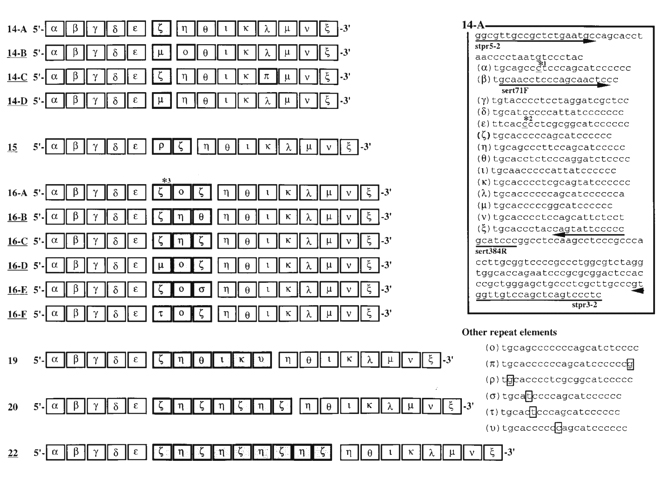
Nakamura et al. 2000
Budget
In development
Bibliography / References
Rule et al. 1996
Gerlerntner 1993
Mellman 1996
Zhang et al. 2000
Nakamura et al. 2000